Role of Interleukin 10 in the modulation of the airway immune response during lung bacterial infection / |c Hernán F Peñaloza Cerda ; thesis advisor Susan Bueno
Texto completo
(2) 2 INDEX ITEM. PAGES. FIGURE INDEX………………………………………………………………………………... 6. TABLE INDEX…………………………………………………………………………………... 9. SUMMARY……………………………………………………………………………………... 10. RESUMEN……………………………………………………………………………………... 12. INTRODUCTION…………………………………………………………………………. 15. 1.. 1.1. Pneumonia……………………………………………………………………... 15. 1.2. Klebsiella pneumoniae………………………………………………………... 18. 1.2.1 Hypervirulent Klebsiella pneumoniae strains………………………….. 18. 1.2.2 Carbapenem resistant and Klebsiella pneumoniae Sequence Type 258…………………………………………………………………………. 19. 1.2.3 Klebsiella pneumoniae virulence factors and immune. 1.3. response evasion……………………………………………………………….. 20. 1.2.4 Immune response agains K. pneumoniae in the airways……………. 24. Streptococcus pneumoniae……………………………………………………. 27. 1.3.1 Implications of antibiotic treatment and polyconjugate vaccine implementation: Emergence of antibiotic resistance and. 1.4. serotype replacement……………………………………………….…. 27. 1.3.2 Immune response against Streptococcus pneumoniae……………... 28. 1.3.3 Bacterial strategies for immune response evasion…………………... 30. Anti-inflammatory myeloid cells: Myeloid derived suppressor cells (MDSCs)…………………………………………………………………... 33. 1.4.1 MDSCs origin and immunosuppressive mechanisms……………….. 33. 1.4.2 MDSCs during bacterial infections……………………………………... 35.
(3) 3 1.5.. Interleukin-10 (IL-10)…………………………………………………………... 38. 1.5.1 IL-10 production and regulation by myeloid cells……………………. 38. 1.5.2 IL-10 effector pathway..….………………………………………………. 41. 1.5.3 The role of IL-10 production during respiratory bacterial infections... 42. 2.. HYPOTHESES………………………………………………………………………….. 46. 3.. AIMS……………………………………………………………………………………... 47. 4.. MATERIAL AND METHODS…………………………………………………………... 48. 4.1 Mice models…………………………………………….…………………………... 48. 4.2 General material and methods: Host/pathogen interaction during Klebsiella pneumoniae Sequence Type 258 infection………………………………. 49. 4.3 General material and methods: Host/pathogen interaction during. 5.. Streptococcus pneumoniae infection…………………………………………………. 62. RESULTS………………………………………………………………………………... 70. 5.1 Klebsiella pneumoniae Sequence Type 258 has acquired resistance to innate immune response……………………………………………………………. 70. 5.1.1 KP35 resistance to neutrophil mediated killing……………………….. 70. 5.1.2 KP35 Resistance to Bone Marrow derived inflammatory monocytes……………………………………………………………….. 75. 5.1.3 KP35 interaction with anti-inflammatory Bone Marrow MDSCs…….. 77. 5.2 IL-10 production by MDSCs is required for host survival during Klebsiella pneumoniae sequence type 258 infection…………………………………………... 79. 5.2.1 MDSCs, neutrophils, eosinophils and Ly6C- monocytes produce IL-10 during KP35.……………………………………………………………... 81. 5.2.2 IL-10 production is critical for KP35 clearance and host survival…... 84. 5.2.3 IL-10 production reduces lung damage during KP35 infection…….. 86. 5.2.4 IL-10 production has a major effect on pro-inflammatory.
(4) 4 cytokine production but does not mostly affect cellular recruitment to the lung tissue………………………………………………………………. 88. 5.2.5 Bone marrow derived MDSCs transferred into IL-10-/- receptor mice restore the resistant phenotype observed in WT mice………………. 93. 5.3 IL-10 production is required for host survival during Streptococcus pneumoniae infection…………………………………………………………. 100. 5.3.1 IL-10 production is necessary for host survival………………………. 100. 5.3.2 IL-10 production impairs bacterial clearance but modulate lung damage…………………………………………………………………….. 104. 5.3.3 IL-10 production modulates neutrophil recruitment to the lungs……. 110. 5.3.4 TNF-α neutralization does not affect the susceptibility observed in IL-10-/- mice……………………………………………………………………... 114. 5.3.5 Two populations of neutrophils are present in the lungs which differs in size and granularity…………………………………………………. 116. 5.3.6 N2 neutrophils are larger and present more granularity as compared to N1 neutrophils…………………………………………………... 118. 5.3.7 IL-10 production during Streptococcus pneumoniae infection does not affect neutrophil activation………………………………………….. 120. 5.3.8 N2 neutrophils are active IL-10 producing cells during Streptococcus pneumoniae infection……………………………………….. 6.DISCUSSION…………………………………………………………………………………. 122 125. 6.1 Klebsiella pneumoniae Sequence Type 258 resistance to primary neutrophils, bone marrow derived monocytes and bone marrow derived MDSCs in vitro………………………………………………………………………….. 126. 6.2 IL-10 production by MDSCs is critical to modulates lung inflammation, KP35 clearance and host survival……………………………………………………. 129.
(5) 5 6.3 IL-10 production during S. pneumoniae infection improves host survival and reduces lung injury………………………………………………………………... 133. 6.4 Identifying and characterizing the heterogeneous population of neutrophils in the lungs of mice during S. pneumoniae infection and the modulatory role of IL-10………………………………………………………………………..……. 135. 6.5 IL-10 production in human respiratory tract infections………………………... 137. 6.6 Concluding remarks: Role of IL-10 production during KP35 and S. pneumoniae pneumonia: Similarities and differences……………………………... 137. 7. ABBREVIATIONS…………………………………………………………………………... 141. 8. SUMMARY OF PUBLICATIONS………………………………………………………….. 149. 9. AKNOWLEDGEMENTS……………………………………………………………………. 152. 10. REFERENCES……………………………………………………………………………. 156.
(6) 6 FIGURE INDEX. Figure number. Figure Tittle. Page. 1. Major characteristics of K. pneumoniae and S. pneumoniae….. 17. 2. Major K. pneumoniae virulence factors…………………………... 23. 3. Immune response against K. pneumoniae………………………. 26. 4. Immune response against S. pneumoniae………………………. 29. 5. Major S. pneumoniae virulence factors………………………….. 32. 6. MDSCs origin and suppressive mechanisms……………………. 34. 7. IL-10 production and regulation…………………………………... 39. 8. Gating strategy for cellular identification during K. pneumoniae infection……………………………………………. 9. Gating strategy for cellular identification during S. pneumoniae infection……………………………………………………………... 10. 74. KP35 resists the clearance mediated by bone marrow-derived monocytes…………………………………………………………... 13. 72. KP35 does not prevent uptake but inhibits phagosome acidification in neutrophils…………………………………………. 12. 64. KP35 infected Neutrophils present an impaired Ca2+ flux ability and an impaired antibacterial function……………………………. 11. 58. 76. BM-MDSCs do not clear K. pneumoniae and express antiinflammatory markers able to suppress neutrophil function……. 78. 14. VertX IL-10/GFP mice pathology during KP35 infection………... 80. 15. Neutrophils, M-MDSCs and Ly6C- monocytes are main IL-10 producers during KP35 infection…………………………... 16. Eosinophils, alveolar macrophages and interstitial. 82.
(7) 7 macrophages are secondary IL-10 producers during KP35 infection……………………………………………………………. 17. IL-10 production is required for KP35 clearance and host survival………………………………………………………………... 18. 95. IL-10 produced by BM-MDSCs is critical for host survival during KP35 pneumonia……………………………………………. 25. 94. Bone marrow-derived MDSCs from VertX and IL-10-/- mice are unable to clear KP35…………………………………………. 24. 92. Bone marrow-derived MDSCs from VertX and IL-10-/- mice present typical MDSCs surface markers………………………….. 23. 91. IL-10 production does not influence immune cell recruitment to BALF of KP35 infected mice…………………………………….. 22. 89. IL-10 production does not influence immune cell recruitment to the lung tissue of KP35 infected mice………………………….. 21. 87. IL-10 modulates pro-inflammatory cytokines production during KP35 pneumonia…………………………………………………….. 20. 85. IL-10 production modulates lung damage during KP35 infection………………………………………………………………. 19. 83. 96. IL-10 produced by BM-MDSCs is critical for restoring bactericidal capacity in IL-10-/- mice but not to reduce pro-inflammatory cytokine production……………………………... 26. The transfer of BM-MDSCs able to produce IL-10 does not affect the cellular recruitment or lung damage of IL-10-/- mice…. 27. 99. S. pneumoniae infection induces IL-10 production in lungs at 48 hpi………………………………………………………………. 28. 98. IL-10 absence lead to a more severe S. pneumoniae. 101.
(8) 8 infection in a dose dependent manner…………………………… 29. IL-10 production facilitates S. pneumoniae dissemination after infection………………………………………………………. 30. 107. IL-10 production modulates the production of pro-inflammatory cytokines during S. pneumoniae infection………………………... 32. 105. IL-10 production modulates lung damage during S. pneumoniae infection………………………………………………. 31. 103. 109. During pneumococcal pneumonia, the production of IL-10 is critical to modulate the amount of neutrophils in the lung tissue but not in the BALF………………………………………………….. 33. IL-10 production does not affect the infiltration pattern of T cells in the lung tissue or the BALF after S. pneumoniae infection….. 34. 111. 113. Etanercept neutralizes TNF-α in vivo but does not improve survival rate of IL-10-/- mice after an infection with S. pneumoniae………………………………………………………….. 35. Two neutrophil subsets, N1 and N2 are found in lungs of WT and IL-10-/- mice during S. pneumoniae infection………………. 36. 121. N2 but not N1 neutrophils produce IL-10 at early time during S. pneumoniae infection……………………………………………. 39. 119. N2 neutrophils from WT and IL-10-/- mice don't show differences in activation during an S. pneumoniae infection……. 38. 117. N2 neutrophils are larger and have more granularity than N1 neutrophils……………………………………………………….. 37. 115. 123. Graphical summary of IL-10 role during K. pneumoniae and S. pneumoniae pneumonia…………………………………………. 140.
(9) 9 TABLE INDEX Table number 1. Table title. Page. Clinical score of mice infected with K. pneumoniae and S. pneumoniae………………………………………………….. 2. 55. Histopathological score of lung tissue during S. pneumoniae infection……………………………………………. 67.
(10) 10 SUMMARY Bacterial pneumonia is a major death cause worldwide, specially in children and the elderly. Two different types of pneumonia have been described: community acquired pneumonia and hospital associated pneumonia. Carbapenem-resistant Klebsiella pneumoniae ST258, an important agent of hospital acquired pneumonia may cause chronic infections with repeated episodes of bacteremia. In this thesis, we evaluated whether a clinical isolate of Carbapenem-resistant Klebsiella pneumoniae ST258 (KP35) is resistant to the clearance mediated by neutrophils and monocytes, which are the most important effectors cells against Klebsiella pneumoniae. During KP35 infection, there is an early recruitment to the lungs of Myeloid-derived suppressor cells (M-MDSCs). In this thesis we provide data that in vitro, BM-MDSCs infected with Klebsiella pneumoniae are highly efficient to suppress neutrophils bactericidal function. We next tested the hypothesis whether the early recruitment of these cells impairs KP35 clearance in vivo. Our data demonstrate early recruited M-MDSCs actively produce IL-10, which, contrary of what initially though, is important for KP35 clearance, reduction of lung damage, cytokine production and improvement of host survival. Bone marrow-derived MDSCs transfer from mice able to produce IL-10 to IL-10-/- mice improved their ability to clear KP35 in the lungs and reversed their susceptibility. On the other side, Streptococcus pneumoniae is a major etiologic agent of community acquired pneumonia worldwide. Some data suggest that the lung inflammation observed during Streptococcus pneumoniae infection facilitates bacterial dissemination. We evaluated the role of IL-10 during pneumococcal pneumonia. As during KP35 infection, IL-10 production improves host survival, modulates the production of pro-inflammatory cytokines and lung injury. However, during Streptococcus pneumoniae, the production of IL-10 led to a reduced bacterial clearance and dissemination and reduced the infiltration of neutrophils to the lung tissue. We next aimed to characterize the infiltrating population of.
(11) 11 neutrophils to the lungs during Streptococcus pneumoniae, we found the existence of two neutrophil subsets in the lung tissue during S. pneumoniae infection in mice, N1 and N2; which have different size, granularity and expression of activation markers. During infection, both neutrophils subsets were increased in the lungs of mice able to produce IL-10, however this increment was significantly increased in the absence of this cytokine. Besides, N2 neutrophils but not N1 neutrophils get activated and produces high amounts of IL-10 during S. pneumoniae. In summary, in this thesis we provide strong evidence about first, the capacity of Carbapenem-resistant Klebsiella pneumoniae ST258 to evade the action of central innate immune response elements; and second about the importance of the early production of IL-10 to allows host survival during bacterial pneumonia caused either with K. pneumoniae and S. pneumoniae..
(12) 12 RESUMEN La producción temprana de IL-10 por células mieloides es un elemento critico de protección del tejido pulmonar durante neumonía de origen bacteriana.. En la actualidad, las neumonías bacterianas son una de las principales causas de muerte en niños y personas de la tercera edad. Dos tipos de neumonía han sido descritas: Aquellas adquiridas en la comunidad y aquellas asociadas a ambientes intra-hospitalarios. Klebsiella pneumoniae resistente a carbapenémicos (typo de secuencia 258) es un importante agente etiológico de neumonía asociada a hospitales. La relevancia de esta bacteria se encuentra en aumento constante debido a su capacidad de establecer infecciones crónicas, las que se caracterizan por repetidos episodios de septicemias. En esta tesis, hemos evaluado la capacidad de KP35, un aislado clínico de K. pneumoniae ST258 resistente a carbapenémicos, de resistir a la actividad anti-bacteriana mediada por neutrófilos y monocitos, los cuales han sido previamente descritos como actores principales en la inmunidad contra K. pneumoniae en el pulmón. Resultados previamente publicados describen el reclutamiento temprano de células Myeloides supresoras monocíticas (M-MDSCs) al tejido pulmonar durante una neumonía causada por KP35. En esta tesis, hemos aportado evidencia concluyente que describe la gran capacidad de MDSCs derivadas de medula ósea (BM-MDSCs) de suprimir la actividad de neutrófilos cuando estos son infectados con K. pneumoniae en condiciones in vitro. A continuación, evaluamos si el reclutamiento temprano de estas células durante una neumonía causada por KP35 impiden la correcta eliminación de esta bacteria en el tejido pulmonar. Nuestros datos, demuestran que M-MDSCs reclutadas el pulmón producen activamente la citoquina anti-inflamatoria Interleuquina-10 (IL-10), la que es necesaria para modular la eliminación de KP35, reducir el daño pulmonar y finalmente permitir la.
(13) 13 supervivencia del hospedero. La transferencia de MDSCs derivadas de médula ósea a ratones incapaces de producir IL-10, los cuales son altamente susceptibles a KP35, logró mejorar la capacidad de estos de eliminar KP35 en el pulmón y a su vez incrementó notablemente la supervivencia de estos ratones. Por otra parte, Streptococcus pneumoniae es uno de los agentes etiológicos más importantes de neumonía adquirida en la comunidad. Distintos artículos científicos sugieren que la alta inflamación y daño pulmonar asociado a la infección facilitan la diseminación bacteriana al torrente sanguíneo y a otros órganos. En esta tesis hemos evaluado el rol de IL-10 durante una neumonía neumocócica. Al igual que durante la infección causada por KP35, la producción de IL-10, modula el daño pulmonar y mejora la supervivencia del hospedero. De manera interesante, durante la ausencia de IL-10 existe una mayor capacidad del hospedero de eliminar S. pneumoniae de los pulmones y evitar su diseminación, lo cual podría estar relacionado con un mayor número de neutrófilos en tejido pulmonar. Dada esta notable diferencia en el número de neutrófilos, procedimos a caracterizar la población de estas células en el pulmón. En esta tesis describimos la existencia de dos sub-poblaciones de neutrófilos, los cuales llamamos N1 y N2. Estas sub-poblaciones se diferencian entre sí en tamaño, granularidad y expresión de marcadores de activación y madurez. Realizamos también una caracterización funcional y evaluamos la capacidad de estas células de producir IL-10. Nuestros resultados muestran un aumento en la producción de IL-10 por parte de neutrófilos totales. Mas aún, esta producción es totalmente aportada por la sub-población N2 y no por la N1. En resumen, en esta tesis aportamos importante evidencia referente, por un lado a la capacidad de K. pneumoniae resistente a carbapenémicos de evadir la eliminación mediada por. elementos clave de la respuesta inmune innata. Por otro lado, nuestros. resultados describen el rol central de la temprana producción de IL-10 por células.
(14) 14 mieloides en la supervivencia del hospedero durante dos modelos de neumonía bacteriana: K. pneumoniae y S. pneumoniae..
(15) 15 1. INTRODUCTION. 1.1. Pneumonia. One of the most severe pulmonary infections is pneumonia. Pneumonia is the second death cause in children under five years old and the elderly worldwide, specially in African, Asian and some Latin American countries [1, 2]. It has been estimated that until 2000 the worldwide rate of pneumonia was 155.8 millions cases per year, concentrated mainly in developing countries with 151.76 millions of cases per year, whereas in developed countries only 4.08 millions of cases per year were registered. From the total cases of pneumonia, 1.9 millions of children died [1]. Two different types of pneumonia can be classified according to where it was contracted: the community acquired pneumonia (CAP) and the hospital associated pneumonia (HAP). CAPs are usually caused by antibiotic sensitive bacteria such as Streptococcus pneumoniae and Haemophilus influenzae type b and typically affects immunocompetent subjects with no other co-morbidities. HAPs, on the other side, are typically caused by antibiotic resistant bacteria, including Pseudomonas aeruginosa, Klebsiella pneumoniae, Acinetobacter baumannii and Staphylococcus aureus. HAPs episodes may be associated with other co-morbilities and immunodepression, but in some cases HAPs are also associated with chronic infections. In this thesis, we will focus on carbapenem-resistant K. pneumoniae (CRKP) and S. pneumoniae, which are very different bacteria but at the same time very important for human health (Figure 1). Whereas CRKP is a major cause of HAPs, two major characteristics make this bacterium a major world concern: the rapid worldwide dissemination and the acquisition of resistance against carbapenems, the last resources used to face this type of bacteria. On the other side, S. pneumoniae is the most important bacterial cause of CAPs. Likewise to CRKP, the increasing antibiotic resistant observed in S. pneumoniae and the lack of effectiveness of current vaccines make this pathogen a major world concern. Due to the lack of.
(16) 16 effectiveness of current therapies against these two major pathogenic bacteria, we believe that the study of the immune response against these two major bacteria is necessary to design new strategies based in the improvement of the immune response. In this context, during this thesis, we studied the modulatory role of the major anti-inflammatory cytokine IL-10 during CRKP and S. pneumoniae pneumonia..
(17) 17. K. pneumoniae. S. pneumoniae. Gram classification. Gram negative. Gram positive. Main diseases in humans. Hospital Associated Pneumonia, Urinary tract infections, Sepsis, chronic infections. Community Acquired Pneumonia, Meningitis, Sepsis, Otitis, Sinusitis. Natural reservoir. Soil, water courses, medical devices, intestinal microbiota. Human nasopharynx. Extracellular or intracellular. Intracellular facultative. Extracellular. Inflammatory response. Weak inflammatory response. Strong inflammatory response. Main innate cells involved in immunity. Neutrophils, monocytes. Neutrophils, monocytes, alveolar macrophages. Figure 1: Main characteristics of K. pneumoniae and S. pneumoniae. K. pneumoniae and S. pneumoniae are very important bacteria in human health although they are very different. K. pneumoniae is a major cause of HAP, it is present in soil, water courses, medical devices and in some cases it can be found in the intestinal microbiota. K. pneumoniae is an intracellular facultative bacterium and despite neutrophils and monocytes are the most important innate cells that respond to K. pneumoniae, the inflammatory response is weak compared to other bacterial infections. On the other hand, S. pneumoniae is a major cause of CAP. It can be found in the nasopharynx as a commensal bacteria. It is an extracellular bacteria and stimulates a strong proinflammatory response mostly driven by neutrophils and monocytes..
(18) 18. 1.2. Klebsiella pneumoniae. Klebsiella pneumoniae (K. pneumoniae) is a Gram-negative encapsulated pathogenic bacterium mostly related to intra-hospitalary infections [3]. K. pneumoniae is an extremely versatile bacterium able to colonize different environments including different kind of soil, water courses, medical devices and mucosal surfaces [3, 4]. This versatility is mainly due to its increased ability to acquire DNA from the environment [5]. In terms of human health, K. pneumoniae is a major pathogenic bacterium, which has had an increased impact over the last 30 years mainly due to two independent phenomena: the emergence and dissemination of antibiotic multi-resistant strains and hypervirulent strains [3-7]. 1.2.1 Hypervirulent Klebsiella pneumoniae Hypervirulent K. pneumoniae (HVKPN) is a highly virulent type of K. pneumoniae able to infect healthy patients in the community, causing severe diseases such as pyogenic liver abscess, meningitis, endophthalmitis and pneumonia; all of them associated with high mortality [6]. HVKPN was first described in Taiwan, and it rapidly became a major concern in Asia [6], although nowadays infections by this bacterium have been reported in USA, Australia and Europeans countries [8-10]. HVKPNs belong to K1 or K2 serotypes and are characterized by a thicker hypermucoviscous capsule and with an incredibly high ability to produces siderophores and incorporate iron [4]. The mucoviscosity-associated gene A (magA), the regulator of mucoid phenotype A (rmpA) and siderophores such as Aerobactin, Enterobacter, Yersiniabactin and Samochellin are responsible of these enhanced virulence observed on HVKPN strains [4]. The traditional treatment against HVKPN is the combined antibiotic administration and drainage of liver abscess. However, the success of the treatment depends in the early.
(19) 19 identification of the infective bacteria [11]. This hard scenario may be even worse if HVKPN acquire resistance to carbapenems. Indeed, last year an outbreak caused by HVKPN resistant to carbapenems was reported in China [7]. The mortality associated was 100% (5/5) and in all cases, the infected bacteria belonged to the sequence type (ST) 11, which presented a hypermucoviscous capsule and resistance to neutrophil-mediated clearance [7]. 1.2.2 Carbapenem resistant Klebsiella pneumoniae Sequence Type 258 In general terms, there are two different types in antimicrobial resistance mechanisms present in K. pneumoniae: the expression of extended-spectrum βlactamases (ESBLs) and the more problematic type, the expression of carbapenemases, which render resistance to virtually all β-lactams, including carbepenems [3]. At least three different carbapenemases have been described around the globe: KPC-β-lactamases (KPCs), Metallo-β-lactamases (MβL’s) and OXA-type β-lactamases (OXAβL’s). MβL’s and OXAβL’s have been mostly confined in Asia and the Middle East/North Africa respectively, although a few cases have been reported in Europe, USA and Latin America [3]. Contrary to MβL’s and OXAβL’s, KPCs have spread all over the world relatively fast [12]. KPCs are only encoded in the Tn4401 transposon, which suggest one or a few original sources of KPC disseminated within and between bacterial species [3]. Tn4401 is highly variable, indeed, eight different Tn4401 isoforms have been described [13]. The differences between each isoform consists in deletions upstream (a,c,d,e,f and n), inside the KPC gene (d) or without any deletion (b) [13]. Multi-drug resistant (MDR) K. pneumoniae is a major cause of HAPs and because is an opportunistic pathogen, infected patients usually present a diminished immune system, unable to clear the bacteria. This phenomenon, combined with reduced possibilities of treatment, results in elevated mortality during infections caused by this pathogen [3]..
(20) 20 However, in rare cases, HAPs caused by MDR K. pneumoniae lead to the establishment of chronic infections characterized by repeated episodes of sepsis [14], which is a remarkably different outcome compared with classical or HVKPN infections. The current classification of K. pneumoniae is based on a multi-locus analysis [4, 5]. K. pneumoniae sequence types (STs) are determined by the nucleotide sequence of 7 loci (gapA, infB, mdh, pgi, phoE, rpoB and tonB). Related STs are grouped in Clonal Complexes (CCs). CC258 is the most widespread Clonal complex worldwide and is composed by ST258, ST11 and ST437 [15]. Among them, ST258 is the most widespread ST [16]. To evaluate whether the worldwide disseminated Carbapenem-resistant K. pneumoniae ST258 (CRKP-ST258) belongs to one single clone or whether was due to multiple divergent events, concatenated single nucleotide polymorphisms (SNPs) phylogenetic analyses were performed, demonstrating the existence of two different clades of CRKP-ST258 with an average difference of 529 SNPs, concentrated in a divergence region of 215-kbps within the capsule-encoding gene island. Interestingly, the two clades presented differences in the wzy gene, which encodes the capsule polymerase [16]. The ratio of nonsynonymous-to-synonymous SNPs indicates that the region of divergence of clade 1 was originated independently of clade 2 [16], suggesting a divergent evolution between each clade. 1.2.3 Klebsiella pneumoniae virulence factors and immune response evasion. As a pathogenic bacterium, K. pneumoniae has several major virulence factors that allow it to evade the host immune response [4, 5]. One of the most studied virulence factor is the Polysacharide Capsule (CPS). CPS is encoded in cps locus, which includes wzi, wza, wzb, wzc, gnd, wca, cpsB, cpsG and galF genes [4]. Genetic regulation over cps locus allows K. pneumoniae a plasticity on CPS synthesis depending on environmental.
(21) 21 conditions; whereas high glucose concentration enhances the CPS production through the activity of RmpA, high extracellular iron concentrations inhibit the CPS production. The CPS mediates resistance against several immune response elements [5]. For example, it mediates resistance against anti-microbial peptides, prevents the binding of antibodies and complement system, inhibits macrophages/neutrophils phagocytosis, participates in biofilm formation and facilitates intestinal colonization [4, 5] (Figure 2A). Importantly, the increased virulence observed in HVKPN is related with a thicker hypermucoviscous CPS and the ability of these strains to block/neutralize the host immune response [4, 11]. Type 1 fimbriae and type 3 fimbriae are other virulence factors that mediate cell adherence of K. pneumoniae [4]. These structures mediate biofilm formation and the adherence to biological and non-biological surfaces, including host cells, the gastrointestinal tract, lungs tissue and urinary catheters [4, 5] (Figure 2B). The Lipopolysaccharide (LPS) is another important virulence factor of K. pneumoniae that contributes to infection. LPS is composed by an intern part, the Lipid A, attached to the external membrane; a medium part, which is the Oligosaccharide core and the external portion, the O antigen [4, 5]. Lipid A and O antigen are major virulence factors that participate in K. pneumoniae immune response evasion (Figure 2C). Despite Lipid A is easily recognized by the immune system (TLR-4), it is also important to provide resistance against antimicrobial peptides [4, 5]. Moreover, modifications on Lipid A composition is related to a diminished recognition by the immune system and a reduction of cytokine production [4, 5]. The O antigen, the LPS external portion, also participates in immune evasion by inhibiting macrophages/neutrophils phagocytosis, inhibiting cytokine production, kidnaping the complement protein C3b and therefore, providing resistance to complement-mediated clearance [4, 5]..
(22) 22 Outer membrane proteins (OMPs) and porins are also important mediators of immune response evasion. OmpA has been one of the most described OMP involved in K. pneumoniae virulence. This protein facilitates adhesion to lung epithelial cells [4, 5]. OmpK35 and OmpK36 porins are important to impair phagocytosis [4, 5], although it has been described a down-regulation of these proteins in antibiotic-resistant strains of K. pneumoniae, perhaps as a strategy to avoid the entry of antibiotics [4, 5]. Efflux pumps, such as AcrAB have been indeed, associated with antibiotic resistance (Figure 2D). Siderophores and iron pumps are important virulence factors (Figure 2E). One major protein involved in the iron transport inside the bacterium is KFU, which absence highly diminished the bacterial invasion ability and virulence [4]. Among the siderophores, four siderophores have been described in K. pneumoniae: Yersiniabactin, Enterobactin, Aerobactin and Salmochelin. HVKPN isolates typically express two or three different siderophores, and its virulence is associated with the production of siderophores [4]..
(23) 23. B. Type I and Typ3 III Fimbriae. A. Polysaccharide Capsule. Biofilm formation Antimicrobial peptides. Phagocytosis. Biofilm formation. Adherence. Type III. Type I. Capsule. C. Lipopolysaccharides (LPS) LPS O-Antigen. Phagocytosis, complement system, cytokine production. Lipid A. Antimicrobial peptides. D. Outer membrane proteins Phagocytosis. OmpK35/36. Adhesion to epithelial cells. Efflux pump; antibiotic resistance. OmpA. AcrAB. E. Siderophores Fe3+. Yersiniabactin, Enterobactin, Aerobactin, Salmochelin. Iron pumps. Figure 2: Major K. pneumoniae virulence factors. K. pneumoniae has several virulence factors that allow it the evasion of the inflammatory immune response. (A) The polyssacharide capsule prevents the action of antibacterial peptides, phagocytosis and promotes biofilm formation. (B) Types I and III fimbriae are required for adherence and biofilm formation. (C) Surface lipopolysaccharides interfere with phagocytosis, cytokine production and impairs the action of complement system and antimicrobial peptides. (D) Different outer membrane proteins impairs phagocytosis and facilitates adhesion and antibiotic resistance. (E) Several siderophores favor bacterial survival inside the host..
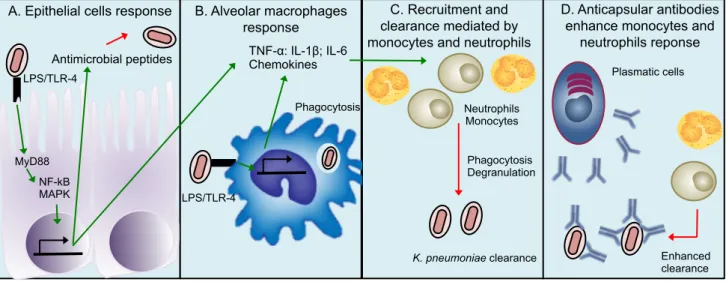
(24) 24 1.2.4 Immune response against Klebsiella pneumoniae in the airways. Once K. pneumoniae colonizes the alveoli, a rapid immune response starts. Alveolar macrophages and epithelial cells are the first defense line against K. pneumoniae in the lungs. Epithelial cells recognize infective bacteria through the binding of bacterial PAMPs to PRRs expressed in their surface, including extracellular and intracellular TLRs, leading to pro-inflammatory cytokines and chemokines production through the pathway TLR/MyD88/NF-𝜅B-MAPK, activating adjacent epithelial cells and recruiting monocytes and neutrophils [17]. These cells also produce antimicrobial peptides such as β-defensins and cathelicidins, which will clear K. pneumoniae. Epithelial cells also secrete Lipocalin-2, which inhibits bacterial growth by eliminating or reducing the ability of K. pneumoniae to kidnap iron [18, 19] (Figure 3A). The recognition of K. pneumoniae by alveolar macrophages, through the engagement of TLRs and NLRs will lead to a phagocytic response that will clear bacteria. These cells also produce pro-inflammatory cytokines and chemokines that will promote the recruitment of neutrophils and monocytes (Figure 3B). When the antimicrobial response mediated by epithelial cells and alveolar macrophages is not enough to clear K. pneumoniae, the recruitment of monocytes and neutrophils becomes critical [20-22] (Figure 3C). The importance of neutrophils in this infection has been assessed using classical antibiotic susceptible K. pneumoniae strains. However, it has been recently described that during HVKPN or CRKP-ST258, neutrophil depletion does not impact significantly in host survival and bacterial clearance, suggesting a secondary role of these cells during infection [23-25]. Ly6C+ Monocytes seems to be critical to clear K. pneumoniae and their depletion led to impaired bacterial clearance, as well as reduced survival [23, 26]. Recruited monocytes during CRKP-ST258 are constituted by a mixed populations of proinflammatory and anti-inflammatory cells. Pro-inflammatory Ly6C+ monocytes express TNF-α and are important to clear K. pneumoniae [26]. Anti-inflammatory monocytes.
(25) 25 express Arginase-1 (Arg-1), iNOS and Interleukin-10, leading to the development of a weak inflammatory response, probably facilitating the establishment of bacterial persistence or at least an impaired clearance [14]. In the bloodstream, there are several antimicrobial elements, including proteins of the complement system and circulating antibodies. Recent reports demonstrate that whereas classical K. pneumoniae strains are resistant to the antimicrobial activity of these components, CRKP-ST258 strains are highly susceptible to the serum [27, 28]. HVKPN, however, maintain the resistance observed in classical strains, apparently due to their hypermucuviscous capsule [27]. The susceptibility observed in different CRKP-ST258 isolates to the serum is due to supportive actions of the innate and adaptive immune components, more specifically the anti-capsule antibodies produced by B cells, which facilitate the binding and microbicidal activity of the complement system [29], as well as bacterial phagocytosis by neutrophils [29] (Figure 3D)..
(26) 26 A. Epithelial cells response. B. Alveolar macrophages response TNF-α: IL-1β; IL-6 Chemokines. Antimicrobial peptides. C. Recruitment and clearance mediated by monocytes and neutrophils. Plasmatic cells. LPS/TLR-4 Phagocytosis. Neutrophils Monocytes Phagocytosis Degranulation. MyD88 NF-kB MAPK. D. Anticapsular antibodies enhance monocytes and neutrophils reponse. LPS/TLR-4. K. pneumoniae clearance. Enhanced clearance. Time Figure 3: Immune response against K. pneumoniae. When K. pneumoniae infects the alveoli, (A) epithelial cells and (B) alveolar macrophages recognize it by TLR4 and produce several chemokines and pro-inflammatory cytokines that promote the (C) recruitment of monocytes and neutrophils that will clear K. pneumoniae. Finally, (D) the antibodies produced by plasmatic cells increase the ability of neutrophils and monocytes to clear K. pneumoniae..
(27) 27 1.3. Streptococcus pneumoniae Streptococcus pneumoniae (S. pneumoniae) is a pathogenic extracellular Gram-. positive bacterium able to colonize and infect the respiratory tract [30, 31]. This bacterium has been described as a major CAPs cause mostly in children under five years old and the elderly [32, 33]. In both groups, most of S. pneumoniae infections occur in developing countries, although pneumococcal diseases also remain a major health problem in developed countries. For example, pneumonia rates in USA for individuals between 65-69 years-old are 18.2/1,000 person-year and 52.3/1,000 person-year for individuals over 85 years-old [34]. 1.3.1 Implications of antibiotic treatment and polyconjugated vaccine implementation: Emergence of antibiotic resistance and serotype replacement. The current strategies used to prevent and treat pneumococcal infections are conjugate vaccines and antibiotics administration. Both approaches have shown only partial effectiveness. The failure of these approaches is thought to be due to two reasons. First, the phenomenon of serotype replacement between serotypes included in the vaccine formulation. This phenomenon occurs due to the increased circulation of those serotypes not included in the vaccine, which increased their incidence and their ability to cause diseases [35, 36]. Three polyconjugated vaccines (PCV) against S. pneumoniae have been globally introduced [37]: PCV7, PCV10 and PCV13. PCV7 was designed to confer protection against 7 serotypes (4, 6B, 9V, 14, 18C, 19F and 23F), but the introduction of this vaccine allowed the emergence of other serotypes of S. pneumoniae not included in PCV7, such as serotypes 1, 5 and 7F (included in PCV10) and 3, 6A and 19A (included in PCV13) [37]. Currently, around 29.4% of the global invasive pneumococcal disease (IPD) cases are caused by serotypes not included in PCV13, being Europe and North America the most dramatic cases with 71.9% and 57.8% respectively [37]. Europe and North America are geographically the zones with most vaccine coverage, suggesting that the.
(28) 28 rapidness of serotype replacement is directly related with the selective pressure caused by the PCV13. Another important phenomenon is the increased spreading of antibiotic resistance genes among different strains. As a consequence of this fact, increased resistance to β-lactams and macrolides have been detected in several countries [38, 39]. 1.3.2 Immune response against Streptococcus pneumoniae The immune response against S. pneumoniae in the alveoli starts with the recognition of bacteria by alveolar macrophages, epithelial cells and soluble components such as antibodies, complement system and surfactant associated proteins. Epithelial cells, as well as alveolar macrophages, express TLRs in their surface that recognize S. pneumoniae. This event triggers cilia movement, β-defensins production, pro-inflammatory cytokines secretion and neutrophil-monocytes chemokines production [31, 40, 41] (Figure 4A). The consequence of these signals is a first neutrophil influx to the lungs (Figure 4B), which constitutes the main host defense mechanism against S. pneumoniae [31, 42]. In parallel, dendritic cells (DCs) migrate to the lymph-nodes to present antigens to T cells and activate them, promoting a synergic Th1 and Th17 response (Figure 4C). This adaptive response induces a second neutrophil-monocyte influx and antibody production by B cells [31, 43, 44] (Figure 4D), resulting in a proper immune response and elimination of pathogenic bacteria from the lungs..
(29) 29 A. Epithelial/ alveolar macrophages response Antimicrobial peptides TNF-α, IL-1β, IL-6 Chemokines. Phagocytosis. PRRs. B. First Recruitment and clearance mediated by monocytes and neutrophils. C. T cell activation by DCs and polarization. D. Second Recruitment and clearance mediated by monocytes and neutrophils. IL-17. Neutrophils Monocytes. TH17. Phagocytosis Degranulation. PRRs. S. pneumoniae clearance. TH1. Enhanced S. pneumoniae clearance. Time Figure 4: Immune response against S. pneumoniae. When S. pneumoniae infects the alveoli, (A) epithelial cells and alveolar macrophages recognize it by PRRs and produce several chemokines and pro-inflammatory cytokines that promote the (B) recruitment of monocytes and neutrophils that will clear K. pneumoniae. In parallel, (C) lung Dendritic cells migrate to the lymph nodes and present antigens to naive T cells, which will acquire a Th1 profile response, stimulating the production of antibodies and Th17 response which will (D) enhance the recruitment of monocytes and neutrophils.
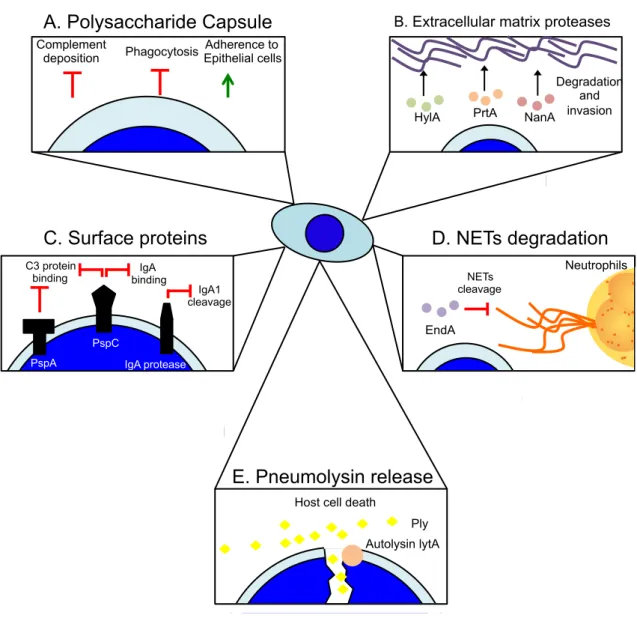
(30) 30. 1.3.3 Bacterial strategies for immune response evasion S. pneumoniae contains several virulence factors used to neutralize the host immune response at different levels. Like to K. pneumoniae, the polysaccharide capsule (CPS) is a major virulence factor of S. pneumoniae [30, 31]. CPS allows S. pneumoniae to escape from the mucus secreted in the whole respiratory tract and once in the alveoli, it protects the bacterium from phagocytosis and complement deposition, and allows bacterial adherence to epithelial cells [30, 31] (Figure 5A). As S. pneumoniae is an extracellular bacterium, it has a strong arsenal of proteins that allow it to neutralize the immune response, to expose surface receptors for adherence and to disseminate within the host. The production of Hyaluronidase (HylA), Neuraminidase A (NanA) and Serin Protease (PrtA), all of them present in the cell wall [31], degrade different components of the extracellular matrix such as hyaluronic acid and N-acetilneuraminic acid, facilitating bacterial invasion [31] (Figure 5B). Furthermore, a wide range of bacterial proteins constitute a complete arsenal of virulence factors used by S. pneumoniae to evade the immune response. Among them, the Pneumococcal Surface Protein A (PspA), which is anchored in the cell wall, has a dual function: on one side binds lactoferrin and apolactoferrin, and by other side prevents the binding of C3 protein from the complement system to the bacterial surface [30, 31]. Pneumococcal Surface Protein C (PspC), another cell wall anchored protein neutralizes the immune response through the impairment of the complement system and the binding to IgA [30, 31]. The inhibition of antibody function by PspC is enhanced by other protein call IgA protease, which effectively degrades the secreted IgA1 [30] (Figure 5C). Finally, the DNase EndA degrades DNA, allowing S. pneumoniae to escape from Neutrophils DNA Extracellular Traps (NETs) [45] (Figure 5D). S. pneumoniae also uses strategies based in.
(31) 31 the induction of immune cell death. For instance, the system LytA/Pneumolysin is the strategy employed to directly kill the immune cells needed to clear S. pneumoniae. The autolisin LytA is a protein required to release Pneumolysin (Ply), which is a cytolytic toxin able to kill leukocytes and epithelial cells (Figure 5E). Ply binds to cholesterol of the lipid membrane and polymerizes, forming pores in the membrane and inducing cell death [30, 31]. Interestingly, the release of Ply also protects viable bacteria, impairing phagocytosis and pro-inflammatory cytokine production by immune cells [46]..
(32) 32. A. Polysaccharide Capsule Complement deposition. B. Extracellular matrix proteases. Adherence to Phagocytosis Epithelial cells. HylA. C. Surface proteins C3 protein binding. IgA binding. PrtA. NanA. Degradation and invasion. D. NETs degradation NETs cleavage. IgA1 cleavage. Neutrophils. EndA PspC PspA. IgA protease. E. Pneumolysin release Host cell death Ply Autolysin lytA. Figure 5: Major S. pneumoniae virulence factors. S. pneumoniae has several virulence factors that allow it the neutralization of the inflammatory immune response. (A) The polysaccharide capsule prevents the action of antibacterial peptides, phagocytosis and promotes bacterial adhesion. (B) The production of extracellular matrix proteases hydrolyzes the extracellular matrix, facilitating the bacterial dissemination within the host. (C) Surface proteins such as PspA, PspC, IgA protease impairs the opsonization mediated by complement system and IgA. (D) The excreted endonuclease EndA cleavages the extracellular DNA, allowing the escape of S. pneumoniae from the action of NETs. (E) The autolysin/pneumolysin system allows the release of the toxin pneumolysin which kills host immune cells..
(33) 33 1.4. Anti-inflammatory myeloid cells: Myeloid derived suppressor cells (MDSCs) 1.4.1 MDSCs origin and immunosuppressive mechanisms MDSCs are cells derived from myeloid progenitors in the bone marrow, released to. the bloodstream [47] (Figure 6). These cells can reach a mature state that suppress different inflammatory processes. How these cells mature is not fully understood, however, it has been described that the environment is crucial. Indeed, several cytokines such as GM-CSF, M-CSF, G-CSF, IL-6, IL-10, and others, have been identified as important mediators of MDSCs maturation [48]. One of the important features of MDSCs is that they are not a homogeneous cellular population. Indeed, two different subsets of MDSCs have been identified in humans and mice: granulocytic MDSCs (G-MDSCs) and monocytic MDSCs (M-MDSCs). In humans, G-MDSCs have been described as CD11b+CD14-CD15+ cells, whereas MMDSCs have been identified as CD11b+CD14+CD16med [49]. In mice, G-MDSCs have been identified as CD11b+Ly6G+Ly6C-, whereas M-MDSCs as CD11b+Ly6G-Ly6C+ [49]. Interestingly, a new MDSCs subset has been recently identified in mice, the EosinophilicMDSCs (Eo-MDSCs) [50]. These cells have been identified during Staphylococcus aureus infection in mice and can be classified as CD11b+Siglec-F+ [50]. G-MDSCs, M-MDSCs and Eo-MDSCs share the same surface markers used typically to identify neutrophils, inflammatory monocytes and eosinophils respectively, making hard to discriminate them from their pro-inflammatory counterparts, especially during infection. For this reason, deeper functional analyses are necessary to the proper identification of MDSCs. These analyses include: the evaluation of their capacity to inhibit T cell antigen-independent and dependent functions, the expression of different transcription factors, the production of cytokines such as IL-10, and transcription of suppressive genes by MDSCs, such as arginase-1 and inducible nos2 (iNOS) [49]..
(34) 34. Inf. Monocytes. Human//mice markers CD11b+CD14+CD16med//CD11b+Ly6C +Ly6G• Arg-1, iNOS, IL-10 producers • Non-Phagocytic. Monocytes. Monocytes M-MDSCs. Immature Neutrophils IMCs. Rb1. Granulocityc precursors G-MDSCs Immature Eosinophils. ? Immature Neutrophils. IMCs Neutrophils. Bone Marrow. Human//mice markers CD11b+CD14+CD16med//CD11b+Ly6C +Ly6G• Pro-inflammatory • Phagocytic • Differentiate into DCs and macrophages. Human//mice markers CD11b+CD14-CD15-//CD11b+Ly6CLy6G+ • Arg-1, IL-10, NOX2/ROS • High MPO • Phagocytic. Human//mice markers CD11b+CD16+CD66b+//CD11b+Ly6CLy6G+ • Highly phagocytic • Pro-inflammatory cytokines • Elastase, MPO, proteases • Antigen presentation • Degranulation. Bloodstream Human//mice markers Unknown//CD11b+Ly6GmedLy6G-SiglecF+ • iNOS. ? Immature Eosinophils. Human//mice markers CD11b+Siglec8+//CD11b+SiglecF+ • Essential to parasitic infections • Highly pathological during asthma and airway allergies. Inflamed tissue. Figure 6: MDSCs origin and suppressive mechanisms. During inflammation, the differentiation of MDSCs starts in the bone marrow, where monocytes, immature neutrophils, immature eosinophils and immature myeloid cells are released to the bloodstream. Once in the bloodstream, these cells arrive to the inflamed tissue and according with the local environment recruited (A) monocytes, (B) immature neutrophils and (C) immature eosinophils may acquire two different phenotype: a pro-inflammatory phenotype and an anti-inflammatory..
(35) 35. Robust data published in the last 7 years support the hypothesis that MDSCs are an immature state of neutrophils [51, 52]. Immature myeloid cells (IMCs) or common myeloid progenitors are the common progenitor between eosinophils, neutrophils, monocytes, DCs, macrophages and MDSCs in the bone marrow [53-55], being the first step in MDSCs development. A report has described that in the EL4 tumor-bearing mice model, M-MDSCs can differentiate and acquire all the morphological characteristics of GMDSCs [52]. This differentiation process is driven by the silencing of the gene encoding Retinoblastoma (Rb1) [52]. Moreover, G-MDSCs share phenotypic and morphological features with neutrophils with some differences, including the expression of some surface markers as CD115 and CD244, reduced expression of the lysosome associated protein LAMP-2, reduced uptake capacity, increased MPO activity and reduced pro-inflammatory cytokine production [51]. Notably, G-MDSCs isolated from tumor-bearing mice can differentiate into neutrophils in culture conditions. After 24 h post-culture, G-MDSCs reach similar uptake capacity, equivalent expression of CD115, CD244 and LAMP-2, and similar capacity to produce TNF-α as compared to neutrophils [51]. These data indicate that in mice, M-MDSCs are G-MDSCs precursors. At the same time, G-MDSCs can potentially differentiate into functional mature neutrophils, suggesting an alternative pathway of neutrophil maturation (Figure 6). 1.4.2 MDSCs during bacterial infections Pathogenic bacteria have developed strategies that allow them to evade the immune response and avoid its clearance. Since MDSCs are able to suppress the immune response, is highly possible that these cells are exploited by bacteria to suppress the immune response. Indeed, the role of MDSCs have been clearly described mainly during chronic.
(36) 36 infections, but also during CRKP-ST258 acute infection. In all cases, infections are caused by major pathogenic bacteria including Staphylococcus aureus (S. aureus), Pseudomonas aeruginosa (P. aeruginosa), Salmonella enterica (S. enterica) and both antibiotic susceptible and resistant strains of Klebsiella pneumoniae (K. pneumoniae). S. aureus is a pathogenic bacterium that usually produces acute infections in several tissues, such as the skin, the airways, gut, bones and soft tissues [56]. However, it has been recently described small colony variants of S. aureus 6850 strain able to cause chronic infections [57]. Mice infected with S. aureus 6850 presented a high bacterial burden in the kidney and in bones after 30 days, and 60% of recovered bacteria corresponded to small colonies S. aureus [57]. In vitro analysis demonstrated a high susceptibility of S. aureus 6850 to macrophages-dependent clearance, indicating that there are host mechanism that are suppressing the inflammatory response against S. aureus 6850 [57]. In agreement with that, a later study demonstrated that during S. aureus 6850 infection, there is a deficiency in T cell proliferation, which was dependent on MDSCs accumulation [58]. S. aureus can accumulate in the form of biofilm in orthopedic implants and other medical devices, causing chronic infections. Around these biofilms an elevated recruitment of CD11b+Ly6C+Ly6G+ MDSCs have been found. These recruited MDSCs are able to inhibit the proliferation of T cells through the expression of iNOS, Arg-1 and IL-10 [59, 60]. Although IL-10 production is not required to inhibit T cell proliferation, it is essential to allow MDSCs recruitment, and bacterial persistence in orthopedic implants [60]. Within soft tissues around the medical device, where S. aureus remains in planktonic form, Arg-1 expression by MDSCs modulates the infection and the bacterial burden [61]. In addition, it has been described that Eo-MDSCs proliferate during chronic S. aureus infection, suppressing the function of T cells by the expression of iNOS [50]. P. aeruginosa is an opportunistic pathogen able to cause chronic infections in the lungs of subjects with cystic fibrosis (CF), chronic obstructive pulmonary disease (COPD).
(37) 37 or critical ventilated patients [62, 63]. It has been demonstrated that CF subjects have increased numbers of G-MDSCs (CD33hiCD66bhiIL-4RαintHLA-DRlow) in the bloodstream as compared to healthy controls [64]. This G-MDSCs accumulation was even higher in CF subjects infected with P. aeruginosa, suggesting the existence of bacterial components that promote the accumulation of G-MDSCs in the blood. Consistently with this idea, human PBMCs cultured in vitro with either flagellated P. aeruginosa or isolated flagellin differentiated into MDSCs, which were able to suppress T cell proliferation [64]. In a mouse model P. aeruginosa also induces the recruitment of T cell suppressive G-MDSCs to the airways [65]. S. enterica is a Gram-negative bacterium able to cause intestinal local infections, such as self-limiting gastroenteritis or systemic infections as typhoid fever in humans [66]. Several serovars of S. enterica have been described, but S. Typhimurium and S. Enteritidis are the most prevalent agents of food-borne gastrointestinal illness [67, 68]. In chronic infection models of S. enterica, the chronic bacterial presence over 90 days in the spleen and liver is correlated with the presence of CD11b+Ly6C+Ly6G- and CD11b+Ly6CLy6G+ cells. These cells have the same surface markers than M-MDSCs and G-MDSCs respectively, however, only CD11b+Ly6C-Ly6G+ cells produce NO and inhibit T cell proliferation and activation, indicating the presence of G-MDSCs and inflammatory monocytes [69]. During antibiotic-sensitive K. pneumoniae infections, MDSCs (CD11b+GR1+) have been described as cells recruited to the lungs at late time post infection to participate in lung recovery, promoting the efferocytosis of apoptotic neutrophils by alveolar macrophages, through the IL-10 production [70]. However, as we discussed above, during antibioticresistant K. pneumoniae infection, M-MDSCs (CD11b+Ly6C+Ly6G-) are recruited to the airways at early time post infection, suppressing the immune response when the bacterium has not been cleared yet [14]..
(38) 38 1.5. Interleukin-10 (IL-10) 1.5.1 IL-10 production and regulation by myeloid cells One of the main suppressive mechanisms employed by MDSCs is the production of. IL-10. Human IL-10 is a 35 kDa homodimeric protein [71] produced by any type of leukocyte depending on the stimulus provided [72, 73]. IL-10 production by myeloid cells starts with the recognition of PAMPs and DAMPs by different PRRs (Figure 7A). In vitro experiments have shown that macrophages and DCs exposed to ligands of TLR3, TLR4 and TLR9 produce IL-10 through two different pathways, involving two different intracellular adaptors, such as MyD88 or TIR-domain-containing adapter-inducing interferon-β (TRIF) [74]. TLR2 stimulation also triggers IL-10 production, since humanderived monocytes treated with Pam3CSK4 - an agonist of TLR2 - also produce IL-10, and the treatment with an anti-TRL2 prevent its production [75]. Similar results were obtained with Pam3CSK4-stimulated murine neutrophils [72]. TLRs are not the only PRRs which induce IL-10 production. PAMPs binding to Ctype lectin receptors also induces IL-10 production in DCs and neutrophils [72, 76]. DCSIGN, a unique C-type lectin present in the surface of macrophages and DCs also induces the production of IL-10 after ligand engagement [77, 78] (Figure 7A)..
(39) 39. A. PAMPs/DAMPs. B. Soluble factors. I. Type I II. Vitamin D III. PGE-2 IFN. Extracellular space Cytoplasm. DC-SIGN C-type Lectin. TLR-2. TLR-4. IFNAR. EP1-4. E. Post-Transcriptional regulation Stimulates Inhibit. MyD88 TRIF. ERK. VDR. MiR-4661. TRAF3. MiR-106q, MiR-98, MiR-27a, TTP. Il-10 translation D. Transcription factors Inhibi Stimulates. C. Epigenetic Metilation. H3. Acetylation. H3; H4. Sp1-3, AP1, STAT3, CREB, C/ EBP, c-MAF, IRFs, NF-kB,Pbx1, Prep1,Atf1, MEF2D, GATA3. t Bcl3. Il-10 il-10 transcription. Nucleus. Figure 7: IL-10 production and regulation. A) PAMPs, DAMPs are recognized by different PRRs present on the cellular surface (DC-sign, C-type lectins, TLR2, TLR4). These receptors induce transcription of the il-10 gene through different pathways that involve either MyD88 or TRIF. B) Soluble factors such as type-I Interferon, prostaglandin (PGE-2) and vitamin D3 also induce IL-10 production. Due to the critical effect of IL-10 on immune cells, several regulatory mechanisms are employed. C) Epigenetic modifications such as H3 and H4 acetylation and H3 methylation influence transcription of the il-10 gene. D) Several transcription factors have been shown to inhibit or stimulate transcription of il-10. E) The translation of Il10 is regulated by the action of microRNAs and by the action of TTP that either inhibit or stimulate IL-10 production.
(40) 40 Co-stimulatory molecules stimulate and reinforce the production of IL-10 after TLRdependent production. TLRs activation in splenic DCs and macrophages stimulates the expression of CD40 in these cells, whose exposure to CD40L in vitro induces the production of IL-10 [74]. Other soluble factors, such as type I IFN, vitamin D3 and prostaglandin-2 have also been identified as important stimuli for IL-10 production [79-86] (Figure 7B). DAMPs are another important IL-10 inductors in leukocytes. Mice exposed to boiling water presented significant production of IL-10 by neutrophils in the spleen [87]. Due to the important modulatory effects of IL-10, its expression must be tightly regulated. Multiple regulatory mechanisms control the production of IL-10, including epigenetic, transcriptional, and post-transcriptional mechanisms (Figure 7C-E). Analyses of DNase hypersensitive sites (HS) on the IL-10 promoter demonstrate the existence of these sites in enhancers and silencers regions. Interestingly, whereas some of these sites are conserved in all leukocytes, others are cell specific [88], suggesting a differential regulation of IL-10 at different cell types. Histone modifications (methylation and acetylation) are also important mechanisms of il-10 regulation. B cells treated with LPS display an increment in acetylation of histones H3 and H4, and methylation of histone H3 in the -1087 region of the il-10 promoter, which activate transcription of il-10 [89]. In monocytes, but not in neutrophils, the acetylation of H3 and H4 and the methylation of H3 allow the binding of different transcription factors involved in the production of IL-10 [90]. The fact that these modifications were not found in neutrophils support the idea of cellspecific regulation. The il-10 gene is also regulated by different transcription factors. Many of these, including specificity protein 1 (Sp1), specificity protein 3 (Sp3), signal transducer and activator of transcription 3 (STAT3), CCAAT-enhancer-binding protein (C/EBP), interferon regulatory factor (IRF), activator protein 1 (AP1), cAMP response element-binding (CREB), the protooncogene c-Maf, nuclear factor-κB (NF-κB), pre-B-cell leukemia transcription factor 1.
(41) 41 (Pbx1), Prep1, cyclic AMP-dependent transcription factor 1 (Atf1), myocyte enhancer factor-2 (MEF2D) and trans-acting T-cell-specific transcription factor (GATA3), positively induce il-10 transcription [84, 91-102]. Bcl-3, in contrast, negatively regulates il-10 transcription [103]. The expression of IL-10 is also regulated at post-transcriptional level. This regulation is mostly accomplished by miRNAs and by the Tristetraprolin (TTP) protein. Specifically, miR-106a [104], miR-98 [105] and miR-27a [106] negatively regulate the translation of IL-10, whereas miR-466l [107] exerts positive regulation. Finally, TTP also negatively regulates the translation of IL-10 [108, 109]. 1.5.2 IL-10 effector pathway The IL-10 receptor (IL-10R) is a complex composed of two different glycoproteins (IL-10R1 and IL-10R2) that belong to the class II cytokine receptor family (CRF2) [110]. The engagement of IL-10 is sequential and involves two steps: first, IL-10 binds to IL-10R1, producing a conformational change of IL-10 that allows it to bind to IL-10R2 [111]. Thus, IL-10 can bind IL-10R1 alone, but not IL-10R2 [112]. IL-10R1 is expressed in all immune cells [113, 114], and in some non-immune cells [115, 116]. Its basal expression is low but is increased after cell activation, for example, neutrophils activated with LPS show an increased expression of IL-10R1 [113]. IL-10R2, instead, is expressed in most cell types and tissues [112, 117], and while it cannot bind to IL-10, it takes part in other receptor complexes that bind other cytokines, such as IL-22 and IL-26 [112, 118]. The signal transduction events developed after binding of IL-10 to its receptor include the receptor-associated Janus kinase family members Jak1 and Tyk2, and the STAT transcription factors STAT3, -1 and -5 [119, 120]. These transcription factors get phosphorylated and associate between them building homo- and heterodimers that.
(42) 42 migrate to the nucleus. At the nucleus, they bind STAT-binding elements in the promoter of IL-10 target genes, inducing their transcription [120]. When IL-10 binds to immune cells, a wild effect of this recognition is the inhibition of proinflammatory cytokines production, reducing the local immunopathology and inflammation [121-126]. However, IL-10 effects are also cell specific, depending on the intrinsic cell function. In macrophages/monocytes, IL-10 enhances phagocytosis, stimulates debris clearance, promotes the anti-inflammatory M2 phenotype expansion and inhibits cellular adhesion to endothelial cells [95, 127-132]. In DCs, IL-10 inhibits maturation, antigen presentation and T cell activation [122, 133, 134]. In B cells, IL-10 stimulates isotype switch and prolong the survival of these cells [135-138]. In endothelial and epithelial cells, IL-10 inhibits the Fas/FasL-dependent apoptosis [116, 139, 140]. 1.5.3 The role of IL-10 production during respiratory bacterial infections The respiratory tract in mammals is anatomically divided into two portions: the upper respiratory tract (nose, pharynx and larynx) and the lower respiratory tract (trachea, bronchia, bronchioles and alveoli). Bacterial infections can affect both portions, although most severe diseases occur during low respiratory tract infections. In these situations, lung failure may be an extremely severe consequence, impairing the air exchange [141]. The role of IL-10 has been studied during infections caused by major chronic pathogens, such as Mycobacterium spp. and Pseudomonas aeruginosa (P. aeruginosa) and has been identified as an important modulator of lung inflammation. M. tuberculosis is the etiologic agent of tuberculosis. This pathogen evades the host immune response and persists within infected individuals for long periods, with most infections deriving from microbial latency. From this state, M. tuberculosis can reactivate and develop secondary disease [142]. The production of IL-10 has been identified as a.
(43) 43 major component of the immunity against M. tuberculosis. In mice, IL-10 neutralization did not affect the survival of mice infected with M. tuberculosis, although it did affect the immune response development, impairing the recruitment of CD4+ and CD8+ T cells to the lungs and diminishing their capacity to produce IFN-γ, thus impacting the host ability to clear the bacterium in the lungs [143]. The cells involved in the production of IL-10 during respiratory M. bovis infection, another relevant Mycobacterium sp., seem to be neutrophils [72]. These cells produce IL-10 and modulate the infiltration of other innate immune cells, such as DCs, monocytes and macrophages; as well as the production of the proinflammatory cytokines TNF-α, IL-1β, and IL-6 [72]. The production of IL-10 by neutrophils during M. bovis is triggered on the one side due to the PRRs recognition of M. bovis [72] and on the other side through the direct neutrophils stimulation by infected DCs [144]. In another study, IL-10-/- mice showed higher susceptibility than WT mice after a low-dose M. tuberculosis infection via aerosol. These animals presented increased lung damage, larger granulomas, higher numbers of infiltrating CD4+ and CD8+ IFN-γ+ T cells and higher amounts of TNF-α, IL-17A, IL-6, G-CSF, GM-CSF and CXCL10 [145, 146], suggesting that the induction of IL-10 by M. tuberculosis is a mechanism exploited by this pathogen to evade the immune response and persist within the host. During chronic P. aeruginosa endobronchial infection, the production of IL-10 is also required to modulate lung damage, pro-inflammatory cytokine production and neutrophil recruitment, but its absence or presence does not impact significantly in host survival or bacterial clearance [147, 148]. The role of IL-10 has also been studied during acute pulmonary infections caused by Francisella tularensis (F. tularensis), a facultative intracellular Gram-negative pathogen able to cause tularemia and pneumonia in humans. During F. tularensis infection, IL-10 production modulates the immune response against this bacterium. The immune response against F. tularensis is mostly Th1 and Th17 [149, 150]. Exacerbated IL-17 production.
Figure


Documento similar
In the preparation of this report, the Venice Commission has relied on the comments of its rapporteurs; its recently adopted Report on Respect for Democracy, Human Rights and the Rule
Therefore, these aspects would confirm that improvements possibly would arise from gains in impulse at swim start obtained specifically on lower limbs with the experimental
Keywords: Metal mining conflicts, political ecology, politics of scale, environmental justice movement, social multi-criteria evaluation, consultations, Latin
Given the much higher efficiencies for solar H 2 -generation from water achieved at tandem PEC/PV devices ( > 10% solar-to-H 2 energy efficiency under simulated sunlight) compared
It is generally believed the recitation of the seven or the ten reciters of the first, second and third century of Islam are valid and the Muslims are allowed to adopt either of
From the phenomenology associated with contexts (C.1), for the statement of task T 1.1 , the future teachers use their knowledge of situations of the personal
Although some public journalism schools aim for greater social diversity in their student selection, as in the case of the Institute of Journalism of Bordeaux, it should be
In the “big picture” perspective of the recent years that we have described in Brazil, Spain, Portugal and Puerto Rico there are some similarities and important differences,





