TítuloParallelizing Epistasis Detection in GWAS on FPGA and GPU Accelerated Computing Systems
Texto completo
Figure


Documento similar
This study, by utilizing genomic convergence of the 12p11 locus with four European GWAS and four gene expression datasets, followed by eQTL and protein network analysis,
candidate gene studies. In other cases, the GWAS approach has identified novel pharmacogenetic associations that would be unlikely to have
Second, SignS is one of the very few genomic analysis tools to use parallel computing. Parallel computing is cru- cial to allow further improvements in user wall time and to
11 Deciphering the role of protein-protein interaction networks in the functional profiling of high-throughput experiments 81 11.1 Ppi networks in Gene Ontology
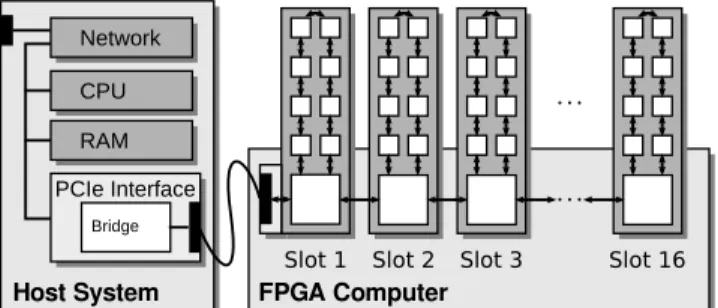
Both FPGA boards can be used also in standard PCs or server machines as endpoint cards in order to provide computational power and/or flexible high
(2013a) Nonequivalent gene expression and copy number alterations in high-grade serous ovarian cancers with BRCA1 and BRCA2 mutations. (2013b) Nonequivalent Gene
(hundreds of kHz). Resolution problems are directly related to the resulting accuracy of the computation, as it was seen in [18], so 32-bit floating point may not be appropriate
Inserting into the flow hash table: The segment data in the GPU segment hash table is inserted in the GPU flow hash table to obtain the number of segments and retransmissions for


