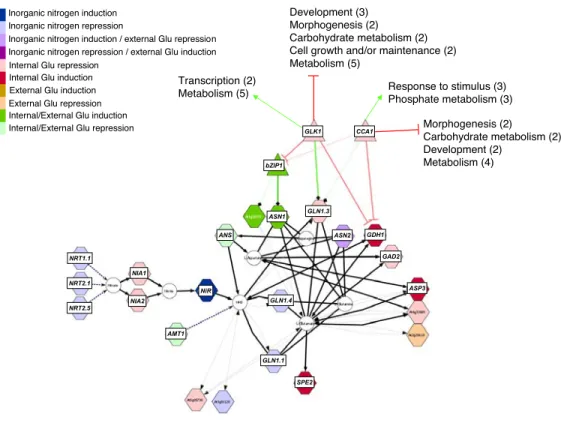
Systems approach identifies an organic nitrogen responsive gene network that is regulated by the master clock control gene CCA1
Texto completo
Figure

Documento similar
We selected 43 genes potentially involved in the regulation of laforin/malin function and/or glycogen metabolism. EPM2A and EPM2B were also included as control of association.
The genome of the soil bacterium Pseudomonas putida KT2440 encodes orthologs of genes crp (cAMP receptor protein), cyaA (adenylate cyclase) and cpdA (cAMP phosphodiesterase) of
Previous work has shown that VEGF up-regulates endothelial Adamts1 expression in a Protein Kinase C (PKC)-dependent manner and has involved the CN-NFAT pathway in Adamts1 gene
Mendelian segregation, the frequency of mutations, and the comprehensive structural and functional analysis of gene variants, identified disease-causing mutations in 12 genes
Importantly, the expression of the FH2 domain, which was mapped as the INF2 domain involved in this process, restored MTOC reorientation and Glu- MT formation in
Diel expression of the circadian clock genes VunGI, VunELF3, VunTOC1, and VunLHY in pod and seed tissue The analysis of the circadian gene network in generative cowpea tissues
This fact favored new species of pollinators such as the hawkmoth that prefers UV- absorbing flowers (white flower) rather than UV-reflecting flowers
The aim of this thesis was to characterise such mechanisms in response to LTD, in order to better understand the regulation of Bdnf gene expression, and its